
| Welcome To Landmark |
Note: This pre-alpha level software is under construction and will change. Not all functionality described is fully implemented or fully debugged. This documentation will undergo revision as the program develops. |
Introduction
Landmark is a
Open2Dprot project Java 2D spot program to Landmark samples into
the system for the Composite
Sample Database (CSD). The CSD consists of protein expression per
(spot) for N samples may be constructred from N sample spot
lists. Landmark is a program to aid in the construction of a
database describing the sample information, region of interest and grayscale
calibration (if applicable for the samples being used).
Landmark is a step [1] module in pipeline analysis for the Open2Dprot project.
You can currently download the pre-alpha version and install it on your computer. Currently, Landmark is hardwired to start with the demo samples and with the -gui switch. The remainder of this home page contains links to some screen shots of the interactive GUI. The Web site contains some initial (rudimentary) documentation.
See the Reference Manual for details. You read about downloading and installing the program on your computer. The source code will be put onto open2dprot.sourceforge.net when it is a bit more stable - currently it undergoing major refactoring.
Please contact us with
suggestions and comments. If you make interesting changes in the
source code, please send us a copy and describe your changes so we can
merge them in the released version.
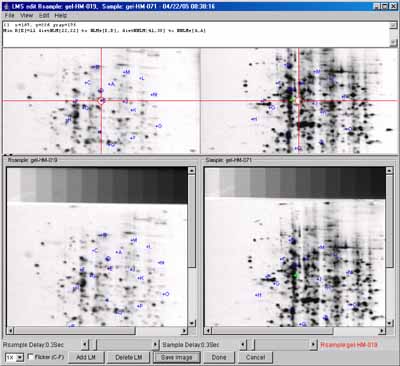
Examples - samples of screen shots
To give the flavor of running the spot pairing, we provide a few
screen shots of the graphical user interfaces and some results. You
can these images in the list below or
view all of the screen shots in a single Web page.
| Contact us | Landmark is a contributed program available at
Open2Dprot.sourceforge.net/Landmark
Powered by |
Revised: 10/04/2005 |